Developing Genomic Resources to Inform Conservation and Management of the Threatened Tricolored blackbird (Agelaius phoeniceus)
Campus: UCLA, Center for Tropical Research
PI(s)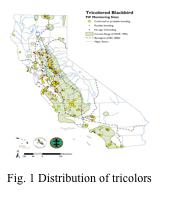 : Kelly Barr, Tom Smith, K. Ruegg and R. Bay
: Kelly Barr, Tom Smith, K. Ruegg and R. Bay
Background: The Tricolored blackbird (Agelaius phoeniceus) is one of California’s most threatened bird species and has been experiencing steep population declines since the 1980’s (Fig.1). At the end of the 19th century, it was described as the most abundant bird species in coastal southern California, but current surveys estimate its population size to be as low as 250,000 with particularly small populations in southern California. Understanding the extent of population structure and connectivity between remaining populations in California is crucial for effective management and conservation, but current genetic resources for this species are limited. Application of conservation genomic tools will allow us to more precisely characterize genetic variation and population genetic structure in this species as well as understand spatial and temporal patterns of movement. These data will provide critical information to help stem further population declines.
Approach: The project will be conducted in two phases. First we will assemble a genome de novo using a combination of Illumina short read sequencing and PacBio Single Molecule Real-Time sequencing. Contiguous sequences (contigs) will be assembled using paired-end 250bp Illumina reads. We have recently implemented this method for both Yellow Warbler and Willow Flycatcher, which resulted in longer contigs (N50=110kb) than the majority of published avian genomes. Contigs will be scaffolded into longer sequences using PacBio reads, which average 10-15 kb. This should result in a high-quality genome that will provide context for future reduced representation sequencing, allowing us to investigate patterns of linkage as well as increasing our power to annotate genes of interest. In the second phase, we will use RAD-Seq mapped to our genome assembly allowing us to genotype over 200 individuals at ~500,000 SNPs. This will provide high-resolution maps of population structure while simultaneously allowing us to infer adaptive processes.
Resources: Samples have been already been collected from all main populations in California. We have one quarter of graduate student support for a graduate student and support for a NSF funded post-doctoral researcher that will assist with running bioinformatics pipelines for genome assembly that has already been developed for the Yellow Warbler and Willow Flycatcher.
